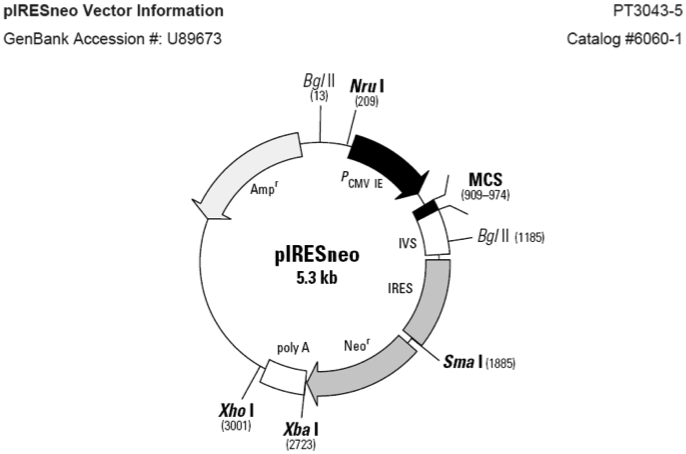
pIRESneo
pIRESneo
编号 | 载体名称 |
北京华越洋生物VECT6064 | pIRESneo |
pIRES-Neo载体基本信息
载体名称: | pIRESneo, pIRES-neo, pIRES neo |
质粒类型: | 哺乳动物细胞表达载体 |
启动子: | CMV IE |
表达水平: | 高 |
克隆方法: | 多克隆位点,限制性内切酶 |
载体大小: | 5.3 kb |
5' 测序引物: | CMV-F |
5' 测序引物序列: | 5'-CGCAAATGGGCGGTAGGCGTG-3' |
3' 测序引物: | pIRESrv |
3' 测序引物序列: | 5'-GCCCTAGATGCATGCTCG-3' |
载体标签: | 无 |
载体抗性: | 氨苄青霉素 |
筛选标记: | 新霉素(Neomycin) |
克隆菌株: | DH5α, HB101 |
宿主细胞(系): | 常规细胞系,293、CV-1、CHO等 |
备注: | 需要将你的基因克隆到 IRES-neo 元件的上游, 本载体已经被Clontech终止销售,被pIRESneo3替代了。 |
稳定性: | 稳表达 或 瞬表达 |
组成型: | 组成型 |
病毒/非病毒: | 非病毒 |
pIRES-Neo载体质粒图谱和多克隆位点信息
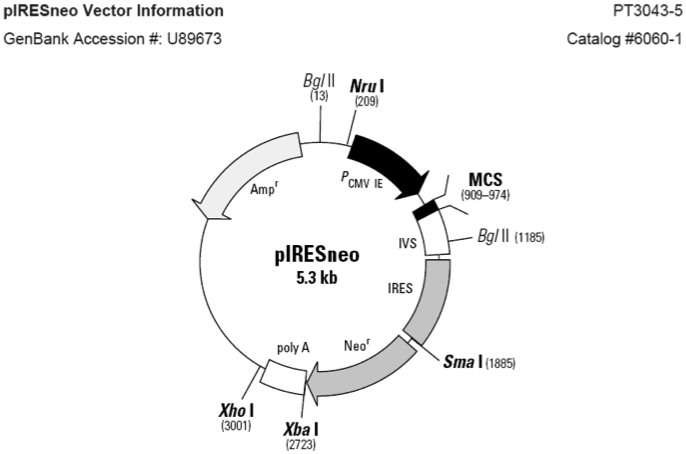
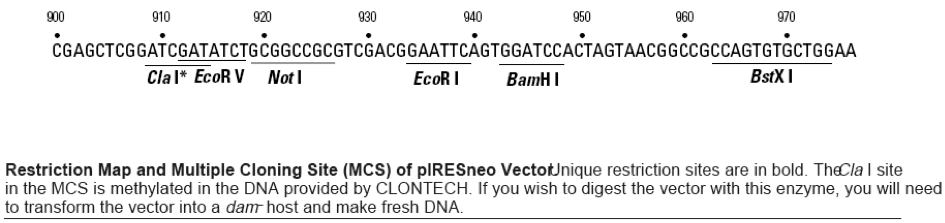

pIRES-Neo载体简介
pIRESneo (originally described as pCIN4 by Rees et al.; 1) contains the internal ribosome entry site (IRES) of the encephalomyocarditis virus (ECMV), which permits the translation of two open reading frames from one messenger RNA (2, 3). After selection with G418, nearly all surviving colonies will stably express the gene of interest, thus decreasing the need to screen large numbers of colonies to find functional clones. The expression cassette of pIRESneo contains the human cytomegalovirus (CMV) major immediate early promoter/enhancer followed by a multiple cloning site (MCS), a synthetic intron known to enhance the stability of the mRNA (4), the ECMV IRES followed by the neomycin phosphotransferase (NPT II) gene, and the polyadenylation signal of the bovine growth hormone. Ribosomes can enter the bicistronic mRNA either at the 5' end to translate the gene of interest or at the ECMV IRES to translate the antibiotic resistance marker.
Use
When using the pIRESneo Vector, the antibiotic exerts selective pressure on the whole expression cassette; thus, a high dose of antibiotic will select only cells expressing a high level of the gene of interest. This selective pressure also ensures that the expression of the gene of interest will be stable over time in culture. Unless your expression experiments require a pure population of cells, you can use the pool of cells surviving selection instead of isolating and characterizing clonal cell lines. We recommend selecting mammalian cultures in 500–1,300 mg/ml of G418 (#8056-1) depending on the cell line (be sure to establish a kill curve for each lot of G418 to determine optimal selection.
其他哺乳动物表达载体: